Slackware and SlackBuilds.org cited in a scientific publication
Posted: 2019-03-15 | Author: slackalaxy | Filed under: academic, slackbuilds | Tags: codeml, evolution, fubar, gnumeric, hyphy, inkscape, meme, msa, pal2nal, paml, PFAM, PRANK, selection, sequence, slac, slackware |2 CommentsThe latest paper from our research group has just been published in Scientific Reports. A big part of the work was done in silico, using different software tools and databases. All bioinformatics software used locally was installed on a Slackware64 system, almost exclusively from the scripts at SlackBuilds.org. With this post, I would like to give my sincere thanks to Slackware and the SlackBuilds.org community.
Slackware has been my distribution since 2005 (it was version 10.1 when I first installed it) and has taught me a lot about Linux. Joining SBo in 2011 introduced me to the basics of scripting and provided a solid quality check of my SlackBuilds. Also, I received a lot of help with problematic scripts from the community, which was especially important between Slackware releases. Now, many of the programs that I maintain at SBo were used in the paper, called “Computational analysis of the evolutionarily conserved Missing In Metastasis/Metastasis Suppressor 1 gene predicts novel interactions, regulatory regions and transcriptional control“.
Below is an example from Figure 3:
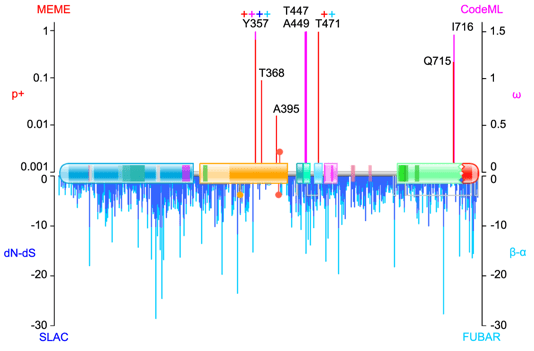
The figure depicts the evolution of a gene, called MTSS1, showing schematically its protein topology with domains and short functional motifs. Protein multiple sequence alignments (MSA) were generated by PRANK for its orthologues from different species. The MSA was then back-translated to codon alignments of the corresponding orthologous CDS (coding DNA sequences) by pal2nal. Selection acting on individual sites (codons and hence the amino acids they code) was estimated by four different evolutionary models/methods. These were SLAC, FUBAR and MEME from HyPhy, and CodeML (Model 8) from PAML. Values were plotted by Gnumeric onto the protein topology scheme, generated by PFAM as described in this earlier post. The figure, as almost all other figures in the paper, was assembled in Inkscape.
Slackware and SlackBuilds.org are credited in “Materials and methods” > “Software and data”. Well… that’s it, I guess :)

Congrats! Score one for the open source community and +5 for the Slackverse.
[…] in Gnumeric, as well as other inconveniences. Like this, I managed to do my work, and in 2019 we published a paper that cordially cites Slackware and […]